What is DNA ligase?
DNA ligase, one of the most-used enzymes in molecular biology, plays a key role in a variety of applications including DNA cloning and next generation sequencing (NGS) library preparation.
DNA ligase is an enzyme that catalyzes the formation of a covalent bond between a 5’ phosphate and a 3’ hydroxyl group at the termini of DNA fragments; this bond ligates the two DNA fragments together. Within cells, these enzymes are essential as they are required for repairing, replicating, and recombining DNA [1]. Because DNA ligases are essential enzymes, they are found throughout the tree of life in archaea, bacteria, and eukaryotes. Despite being widespread, DNA ligases can be largely categorized into just two subgroups—those that rely on nicotinamide adenine dinucleotide (NAD+) and those that adenosine triphosphate (ATP) as a cofactor [1].
How does DNA ligase work?
The mechanism of DNA ligase can be broken down into three steps [2].
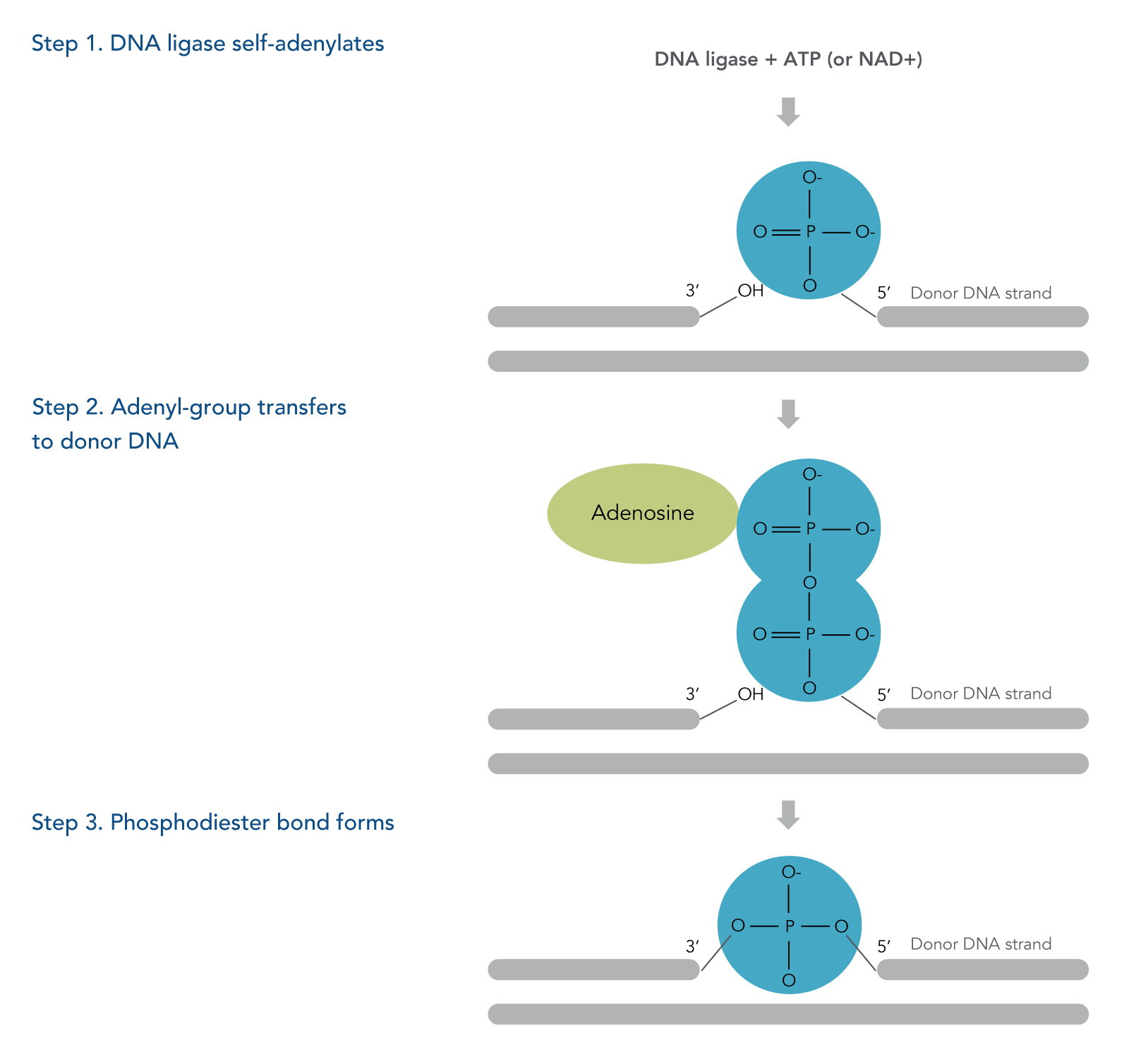
The three steps of a DNA ligation reaction (Figure 1):
- The DNA ligase is self-adenylated by reacting with ATP or NAD+, creating a phosphamide bond between a lysine residue within the enzyme and an AMP molecule of the cofactor, either ATP or NAD+.
- The adenyl group is transferred to 5’ phosphorylated end of the donor DNA strand.
- Phosphodiester bond forms between the hydroxyl group and the DNA adenylate formed in step 2.
Commonly used DNA ligases
Examples of DNA ligases commonly used in molecular biological techniques include the T4 DNA ligase, the E.coli DNA ligase, and thermostable ligases.
E.coli DNA ligase uses NAD+ as a cofactor to bind cohesive double-stranded DNA fragments’ ends. E.coli DNA ligases need unique chemical conditions in order to ligate blunt end DNA fragments, and generally it does not target DNA-RNA or RNA-RNA duplexes efficiently [2].
The T4 DNA ligase originates from bacteriophage T4, and is the most commonly used ligase as it can easily ligate cohesive or blunt ends of DNA and it can reconnect single stranded nicks in DNA, RNA, or DNA/RNA hybrids [2]. Unlike the E.coli DNA ligase, the T4 DNA ligase uses ATP as a cofactor.
As the name implies, thermostable DNA ligases function at higher temperatures than the two previously discussed ligases. This is because they are enzymes that are found in microbes that thrive in extreme temperatures like Thermus thermophilus that can grow in 50–82°C conditions [2,3]. Thermostable DNA ligases are needed for molecular approaches requiring extremely high enzyme fidelity such as ligase chain reaction (LCR) [2].
DNA ligase in action
DNA ligases’ activity depend on a variety of factors such as the concentration of DNA fragments, the ends of the fragments, temperature, etc. It is important to read your DNA ligase protocol and to optimize your ligase reaction according to your unique reaction conditions [2]. As mentioned above, DNA ligases play a key role in DNA cloning and in NGS library preparation.
In cloning applications, DNA ligases are largely used to bond the ends of a DNA fragment to the ends of a plasmid or vector. This vector will ultimately be transformed into cells of interest where it will recombine with the cell’s wild-type DNA, generating a mutant.
For NGS applications, DNA ligases link adapters to DNA fragments, generating a complete library molecule. Adapters are multi-functional in NGS as they facilitate sequencing of DNA fragments by attaching to a flow cell or ensuring capability with a specific sequencing platform. They also contain identifying sequences which allow for multiplexing and sample identification.
You can read more about DNA cloning techniques by downloading IDT’s DNA Cloning Guide here, or to find out more about NGS library preparation download IDT’s Next generation sequencing guide here.